- Superbio AI
- Posts
- 🍬 Superbio Wrapped 2025
🍬 Superbio Wrapped 2025
A year in review: from App Store to Virtual Lab

Dear Superbio community,
2025 was the year Superbio went from useful to transformative, from an AI app store to a full-fledged Virtual Lab.
Together, we ran real experiments, stress-tested new ideas, and pushed Bio-AI workflows beyond demos and into daily research. From first-time users spinning up protocols in minutes, to power users chaining models in ways we never anticipated, this year was defined by momentum.
So before we look ahead, we wanted to pause and reflect on what we built together.
Here’s Superbio Wrapped 2025 👇
🧪 By the Numbers
5,260 model runs executed
900+ computational protocols built
10,000+ GPU hours orchestrated behind the scenes
0 infrastructure setup or coding required 😉
🚀 Top 3 New Features You Loved
Virtual Lab (Cloud Jupyter + GPUs)
End-to-end Bio-AI pipelines, reproducible and runnable in minutes.Plain-English Pipeline Builder
Describe the experiment, Superbio assembles the workflow.Public Superbio APIs
Access and execute 650+ models on Superbio using Python.
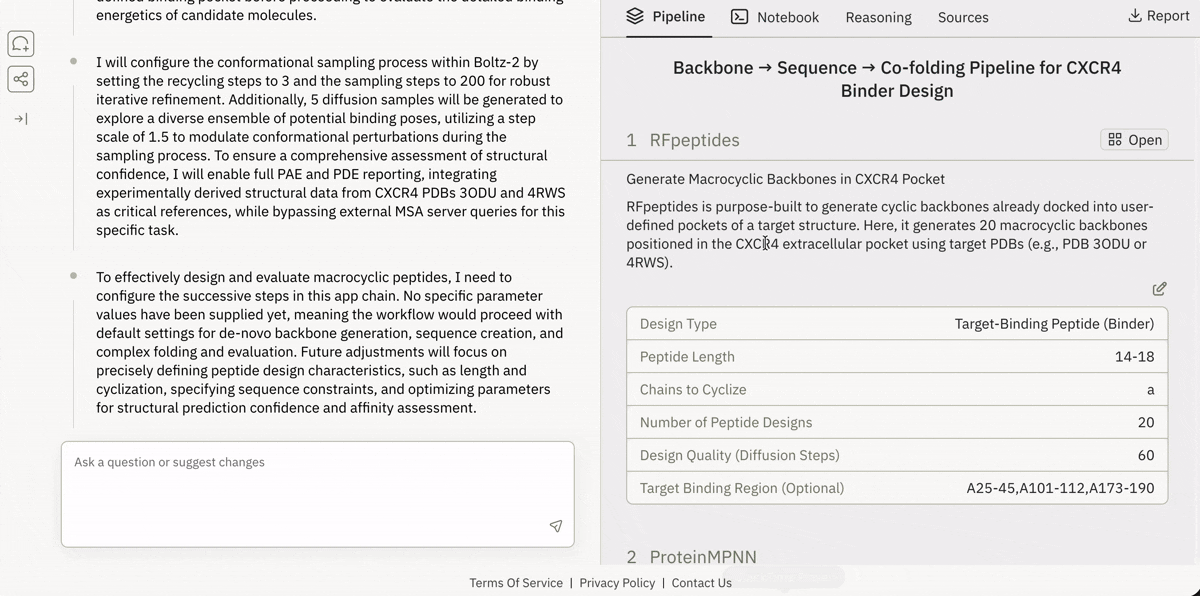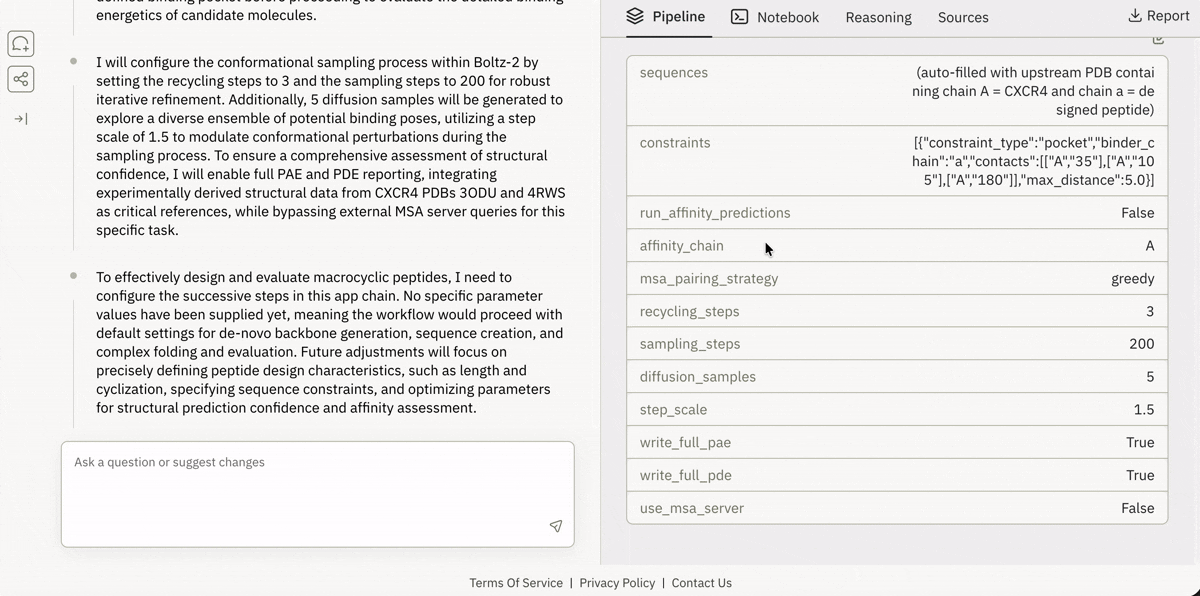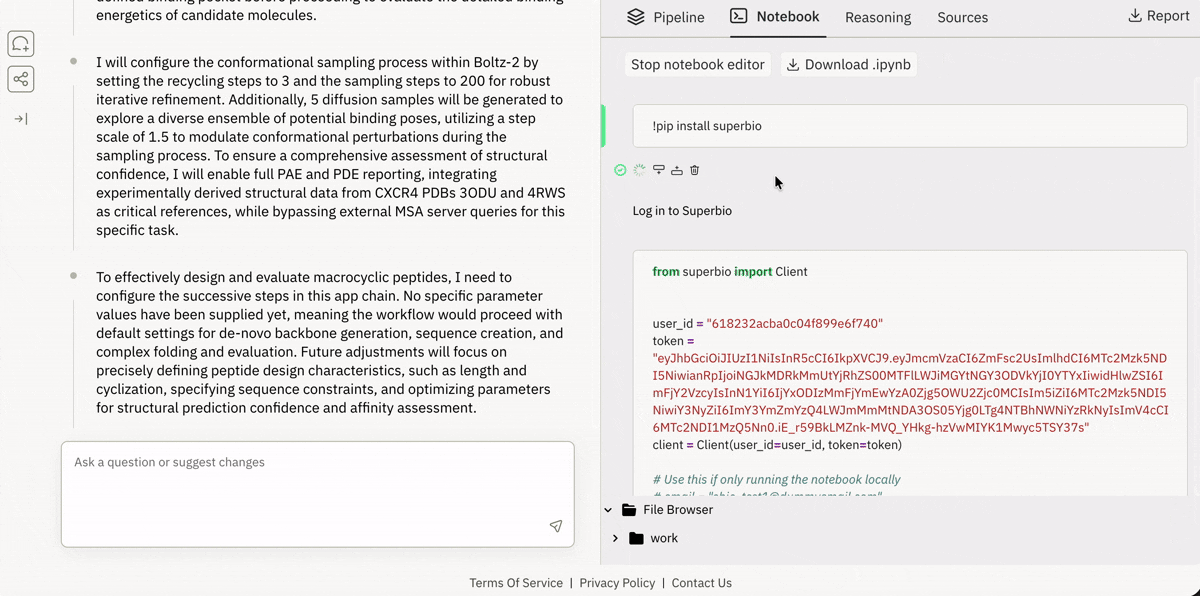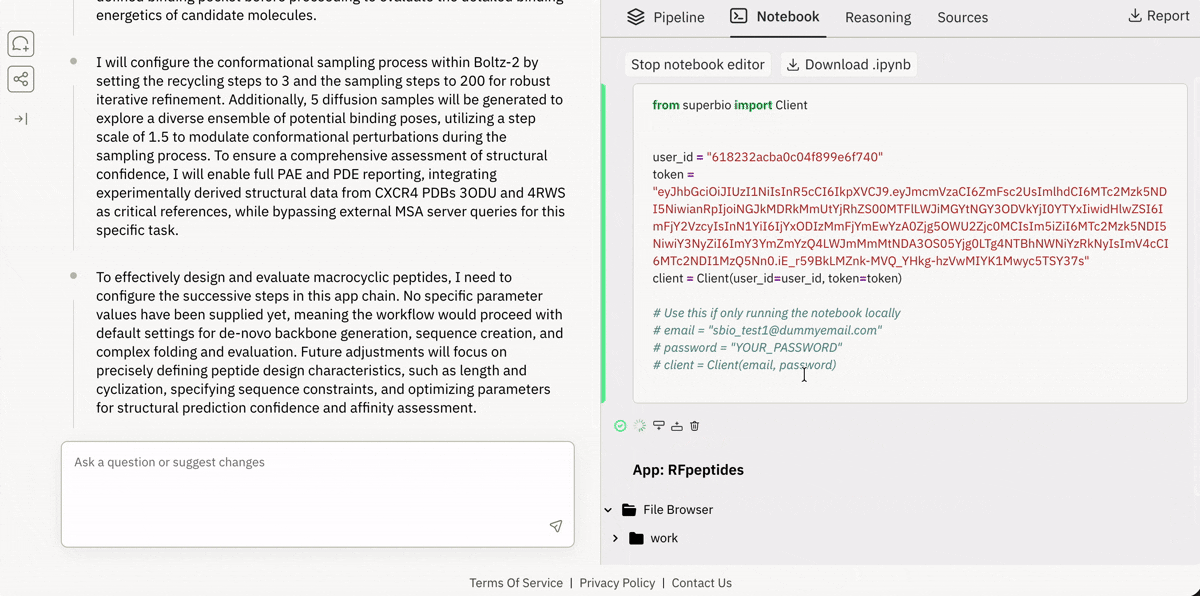
Launching and running a Jupyter notebook in Superbio’s UI.
🧬 Top 3 New Models Added
From generative binder design to structure-aware sequence optimization, this year we expanded our catalog with tools researchers actually use:
Binder Design with Boltz-1 — end-to-end binder design pipeline, scaling to 500,000 binders with ease
LigandMPNN — structure-aware sequence design for protein–ligand and protein–protein interfaces
Boltz-2 — joint modeling of complex biomolecular assemblies and binding affinities
Coming soon: BoltzGen — large-scale generative binder design, built for speed and scale
(What a year for Boltz developers!)
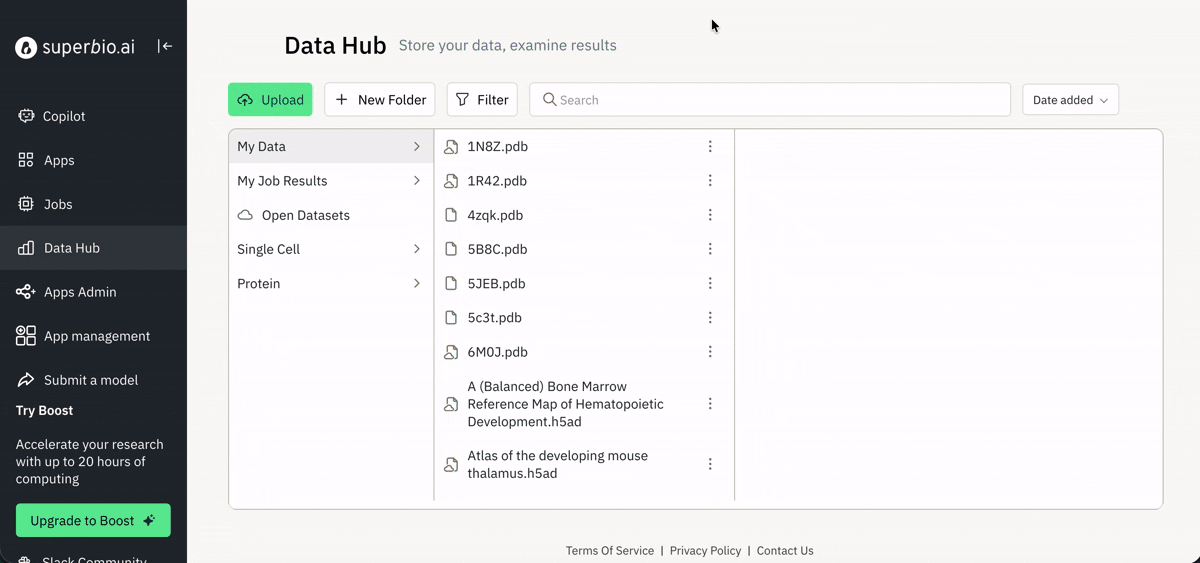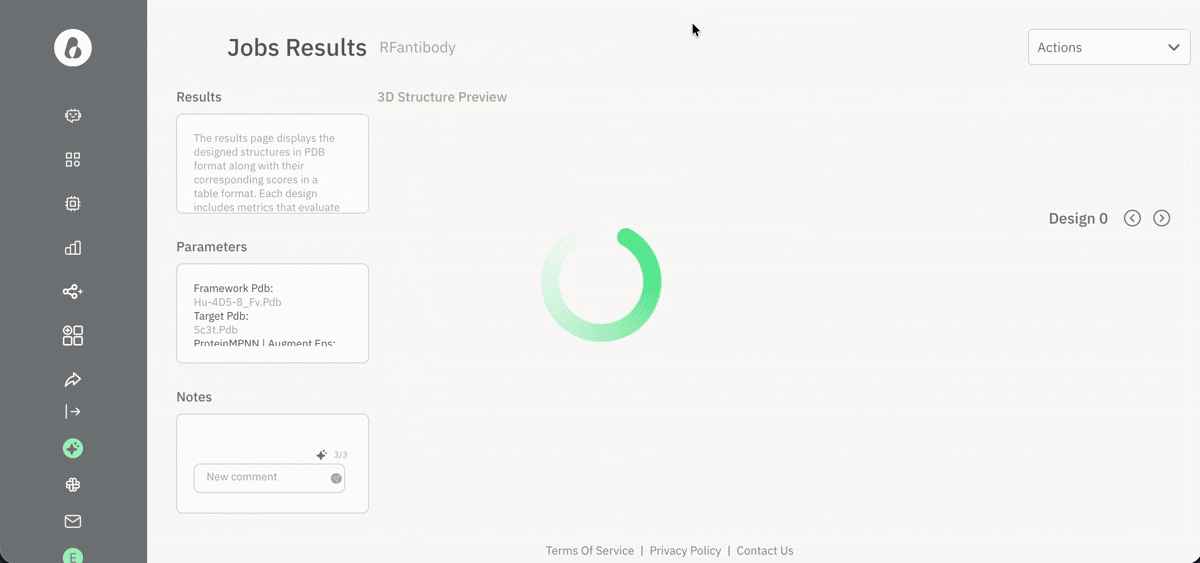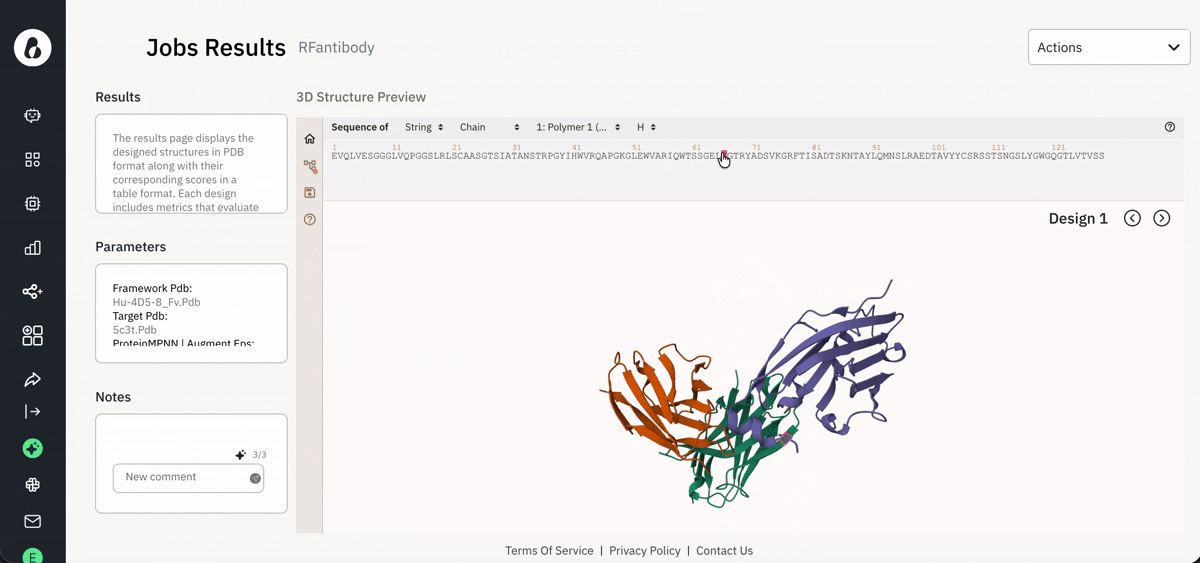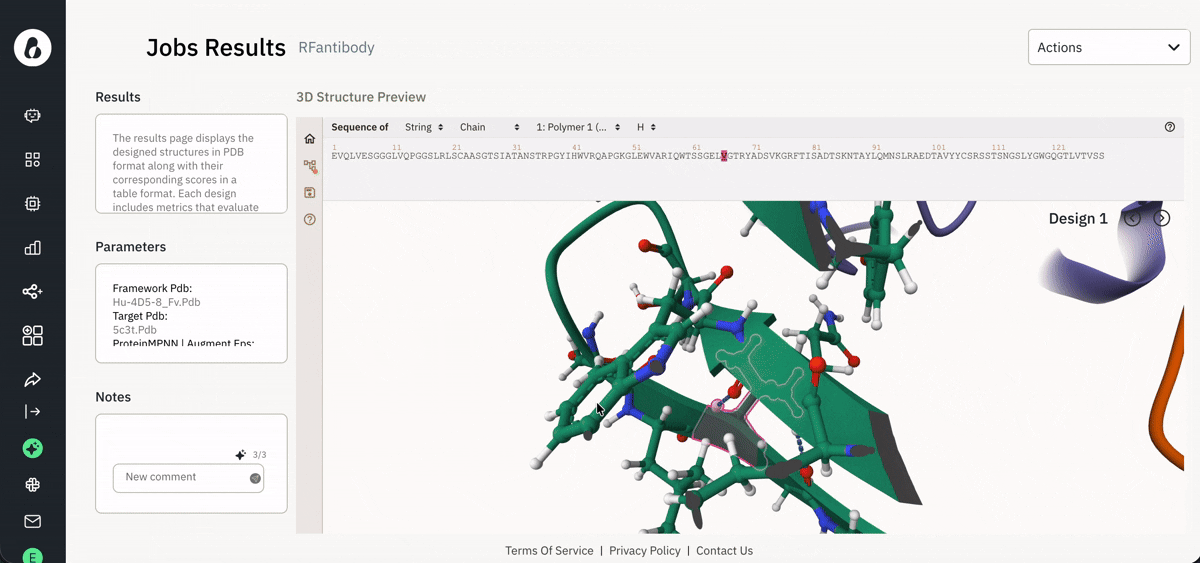
Visualizing results
🧠 Most Common Use Cases
Here’s what researchers used Superbio for most in 2025:
Going from target to real drug candidates
Starting with a biological target and generating binders or variants to exploreScaling exploration before the wet lab
Designing and filtering hundreds of thousands of candidates in silicoAnswering “what should we test next?”
Ranking and narrowing large design sets to a focused shortlistRe-running and sharing experiments
Saving protocols so results are reproducible and easy to revisit
🧑🔬 Community Signals We Loved
Most-requested feature:
An LLM in the loop for interacting with Jupyter notebooks (coming soon 👀)Most common “aha” moment:
Realizing pipelines can be reused, shared, and rerun — not just executed onceMost common user question:
“Can I bring my own models?”
(Yes. Always yes.)
🎓 Top 3 Webinars of 2025
Programming Biology in English — a first look inside Superbio’s Virtual Lab (recording coming soon)
Scientific Agents for Biotech R&D — where we introduced the Superbio Pipeline Builder
Blazing a Trail to Universal Vaccines — a conversation with Dr. Jacob Glanville on the future of vaccines
🔮 Looking Ahead
In 2026, we’re doubling down on:
Enabling full autonomy for the Superbio Co-pilot to execute protocols and help scientists interpret results
Expanding high-quality, production-ready models
Model fine-tuning and head-to-head benchmarking
Thank you for building, experimenting, and pushing Superbio forward with us.
Wishing you happy holidays,
The Superbio Team 🩷