- Superbio AI
- Posts
- Generative AI for Protein Design
Generative AI for Protein Design
Protein-MPNN completes the design loop

Hi all,
Generative AI continues to evolve, offering incredible new tools for the biomedical community - and real opportunities to accelerate rate-limiting steps of the R&D process.
In years past, the process of generating a monoclonal antibody with desired functional properties was anything but trivial. The process involved animal immunization, then cloning a unique white blood cell - an expensive, tedious, and error-prone process. Recent advances in AI/ML have enabled the design of protein binders in silico, reducing this drawn-out discovery process to a matter of days.
Previously, we shared a tutorial leveraging RFDiffusion on Superbio for a hypothetical binder design task targeting human ACE2. Today, we’re excited to share our launch of a second game-changing model on Superbio - Protein-MPNN - enabling us to go from backbone to functional protein structure.
Read on for a detailed how-to of this critical next step 👇
🦴 Generate poly-glycine backbones using RFDiffusion
RFDiffusion, developed by Watson et. al and the Baker lab, offers a revolutionary solution to the protein backbone design problem. It takes advantage of structure-function logic embedded in the RoseTTAFold model, allowing users to specify desired properties of de novo generated proteins.
The Superbio implementation of RFDiffusion allows this to be done without any code. To get started with binder design on Superbio, check out our detailed tutorial here.
🔗 Complete the amino acid chain using Protein-MPNN
RFDiffusion’s function is restricted to backbone-generation - specifically, a poly-glycine sequence with structural information preserved in a PDB file. Therefore, another method must be used to generate functional amino acid sequences.
Enter: Protein-MPNN, a neural network originally developed to scaffold incomplete protein functional sites. Protein-MPNN leverages two computational techniques to achieve this objective: “constrained hallucination” and “inpainting” - techniques which can modify or “fill in” protein sequences while maintaining overall structural integrity.
To get started, simply navigate to Protein-MPNN on Superbio and drop in PDB files of interest.
🧑🔬 Validate polypeptide folding using AlphaFold2
While recent AI/ML advances are amazing, computer-generated polypeptide sequences do not come with functional guarantees.
Before costly lab validation is performed, users can enter amino acid sequences into AlphaFold2 to ensure 3D structure has been preserved via protein sequence generation using RFDiffusion and Protein-MPNN.
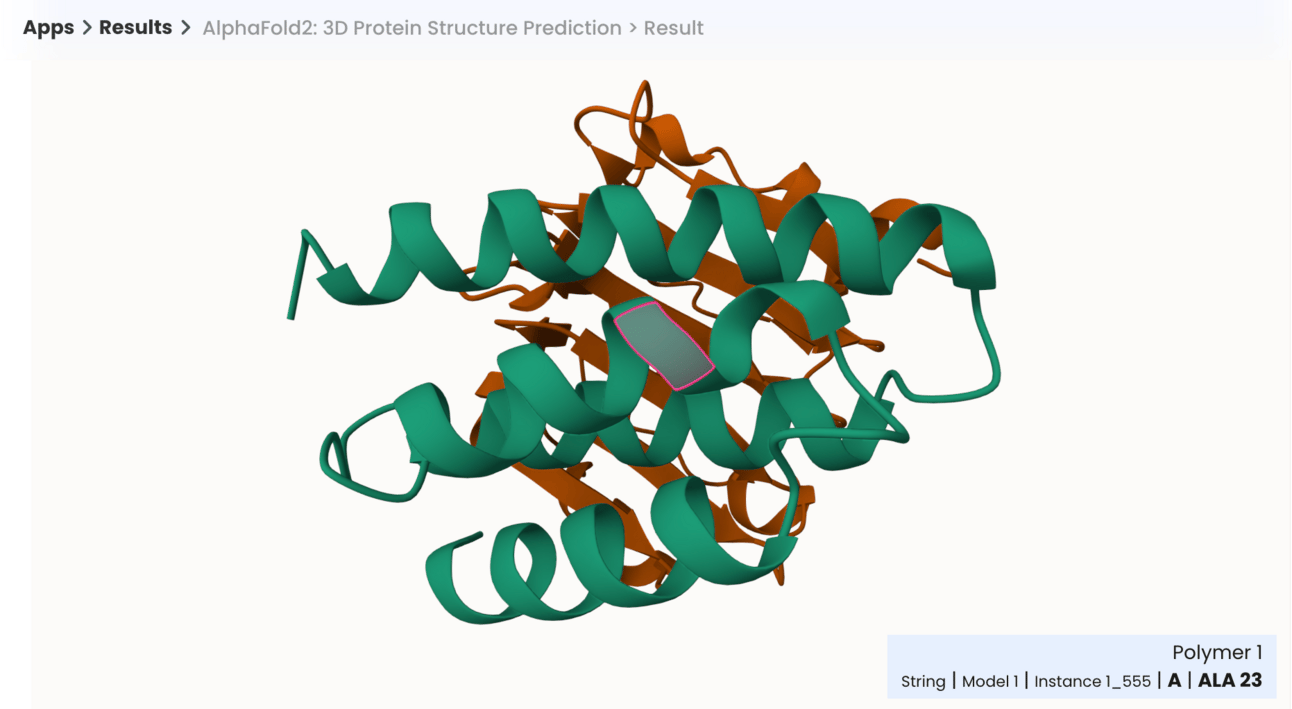

AlphaFold2 results on Superbio, with interactive AA annotations.
Navigate to the jobs page to inspect specific residues, ensuring their expected location in 3D space.
Protein-MPNN has exciting capabilities beyond providing a critical next step downstream of RFDiffusion.
If you’d like to see another tutorial leveraging this tool, feel free to let us know - just hit reply 👉
Stay curious,
Berke from Superbio
P.S. - Want to learn about more of Superbio’s features?
Find our detailed binder design tutorial here.