- Superbio AI
- Posts
- Announcing our “Virtual Lab”
Announcing our “Virtual Lab”
The future of semantic drug design is here.

Dear Superbio community,
Earlier this year, we asked an audacious question: “What if you could program therapeutic design in English?” With the convergence of multiple AI capabilities, our team envisioned that this goal may be within reach for the first time. So, we set out to build the groundwork for an AI-native laboratory - a place where scientists could simply describe a hypothesis, and our system would assemble, execute, and track an experimental workflow from start to finish.
Today, that vision takes a major leap forward. We’re releasing Superbio’s “Virtual Lab” - a product enabling scientists to build and run end-to-end Bio-AI pipelines in a Jupyter environment in the cloud. From binder design to target-ligand affinity predictions, scientists can now design & execute reproducible drug design workflows with plain English - no setup, no infrastructure headaches, no coding experience required.
With this launch, Superbio becomes more than a “Co-Pilot” - it’s evolving into a full-fledged engine for scientific automation. Below are a few capabilities we’re excited to share:
🔗 Spinning up “ready-to-run” drug design pipelines in the browser
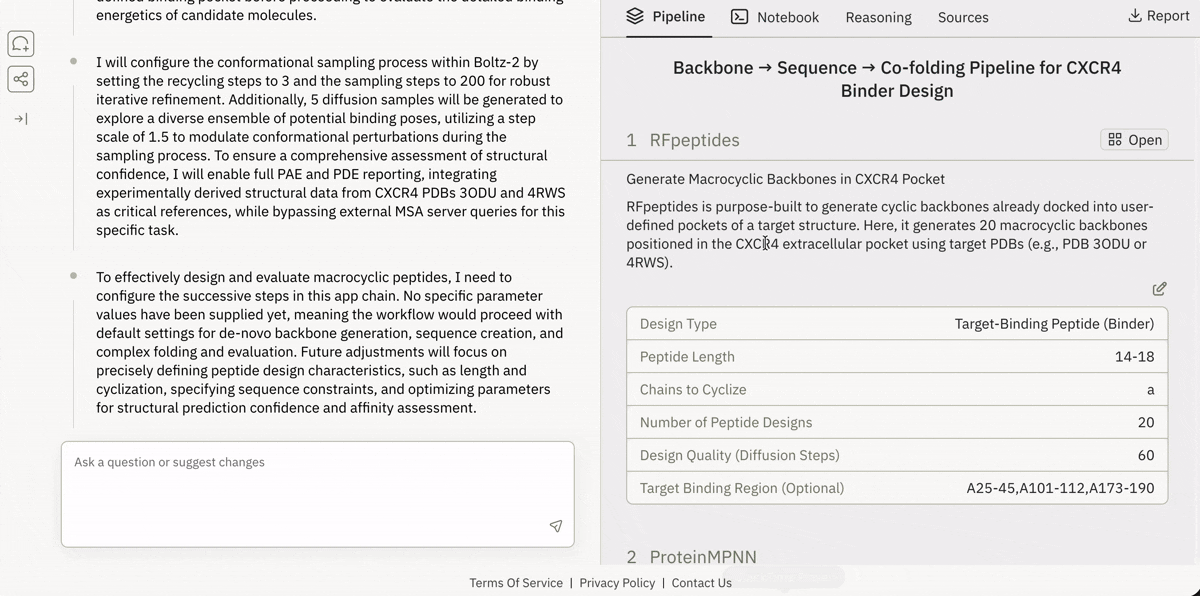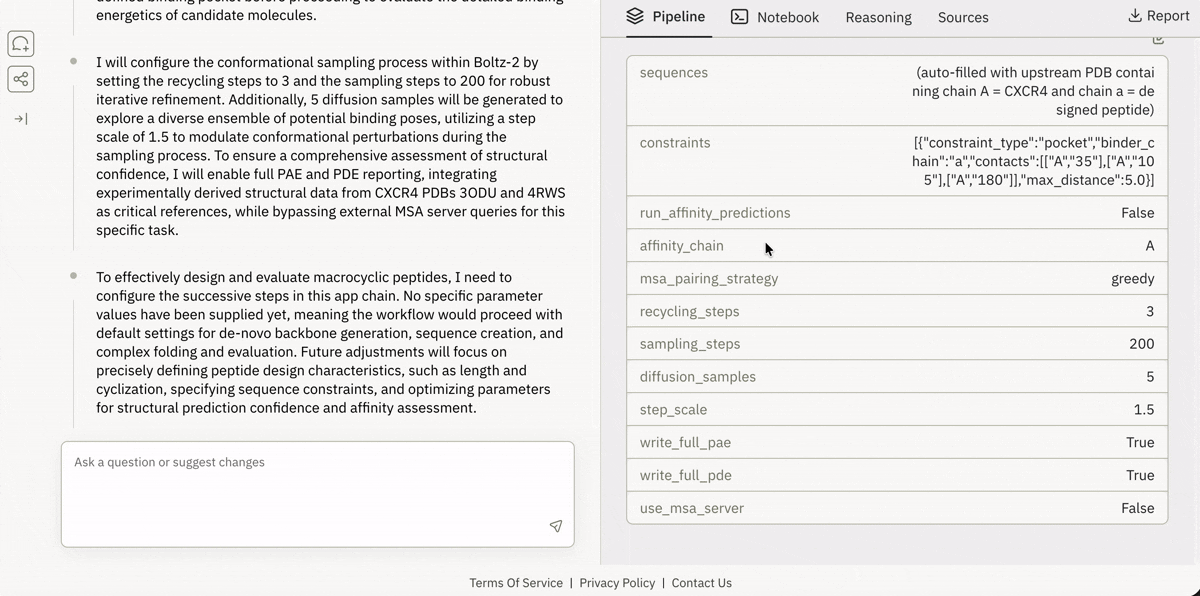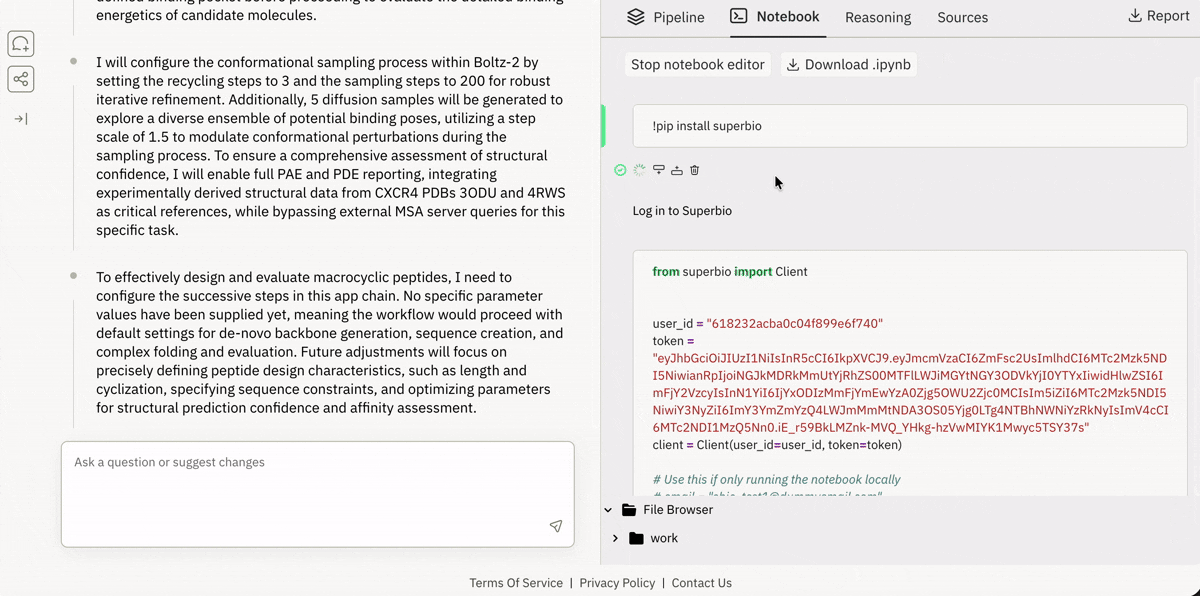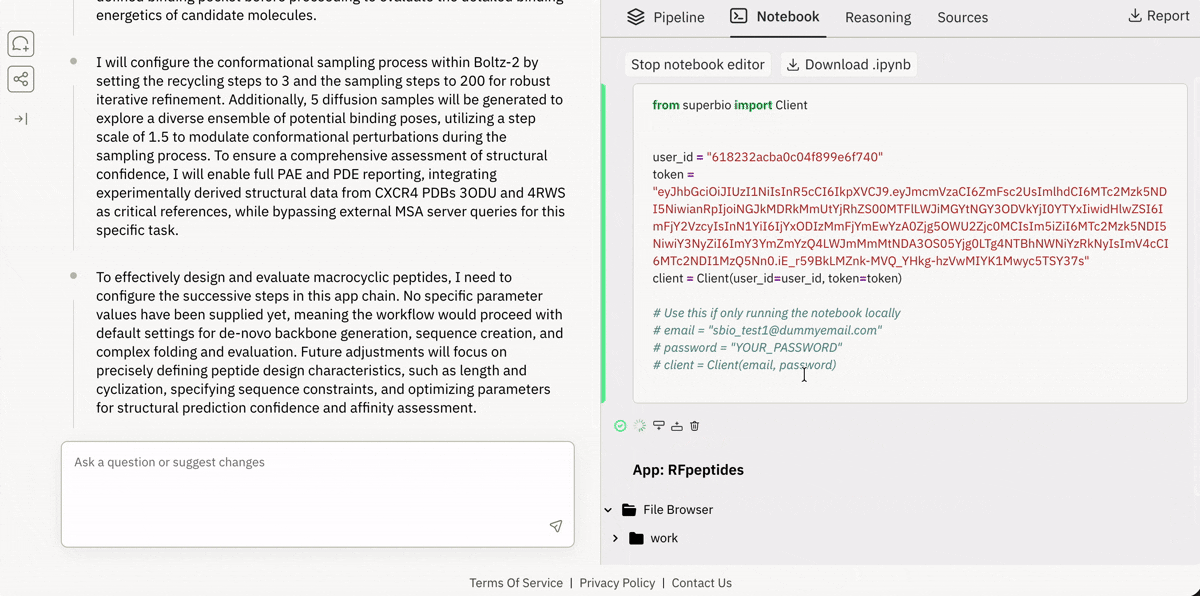
Launching and running a Jupyter notebook in Superbio’s UI.
💾 Incorporating your data from local or remote sources
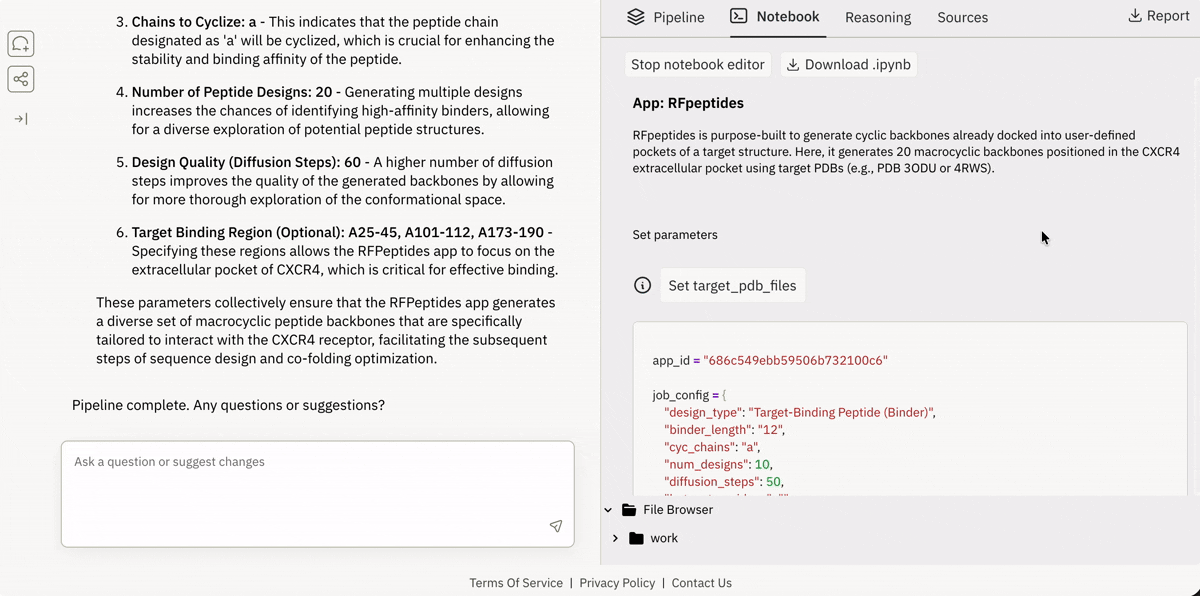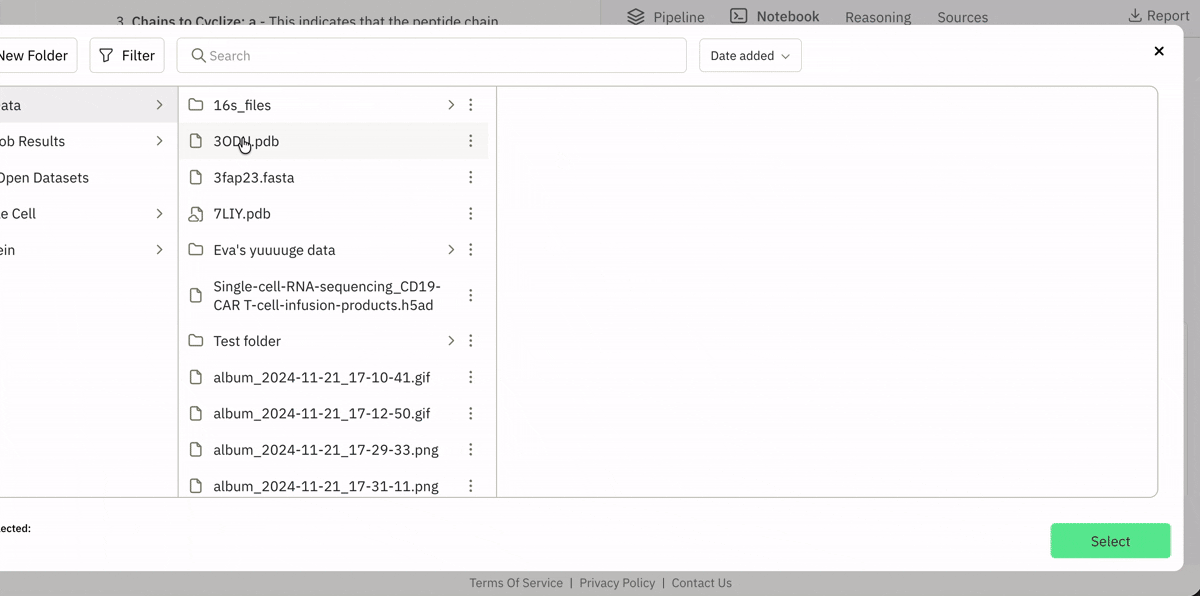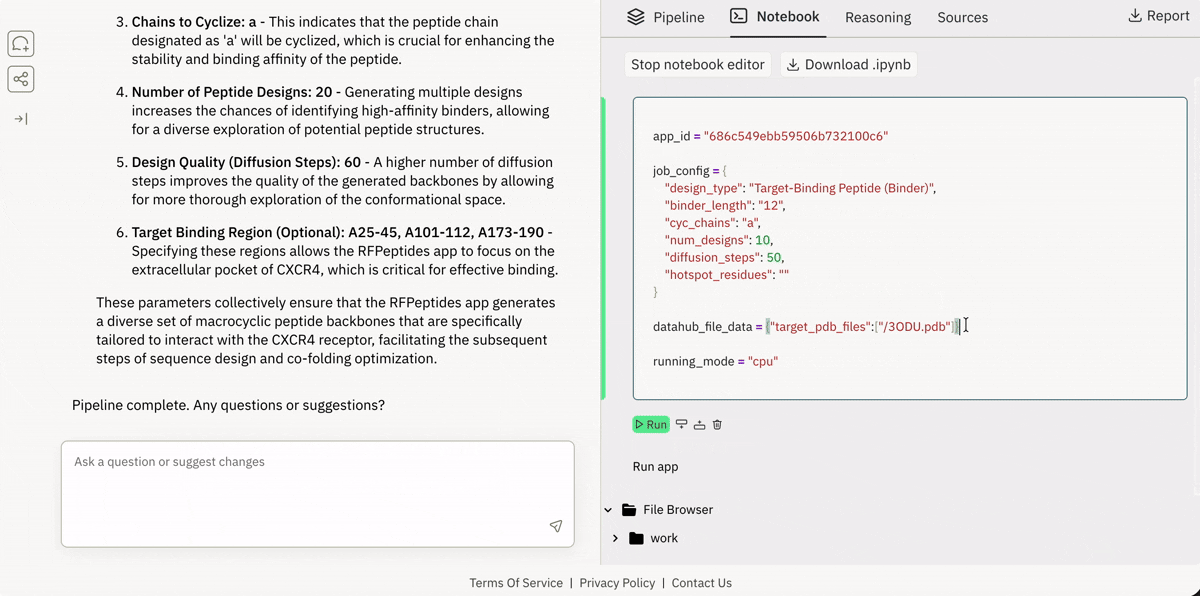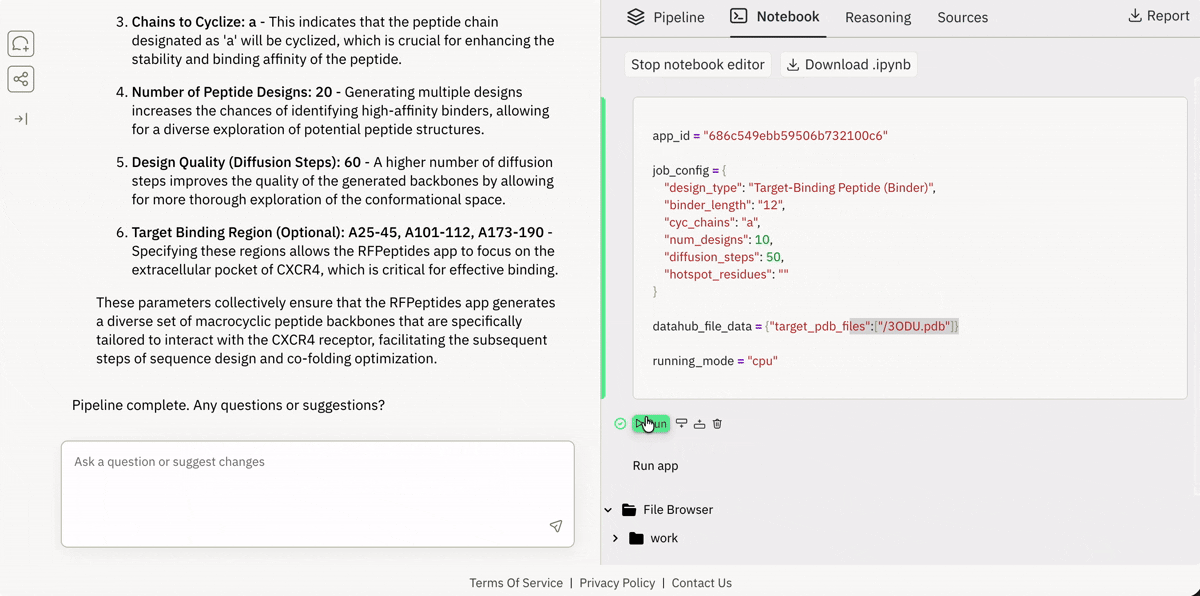
Ingesting data from Superbio’s Data Hub (or other sources).
📈 Viewing and downloading data for analysis
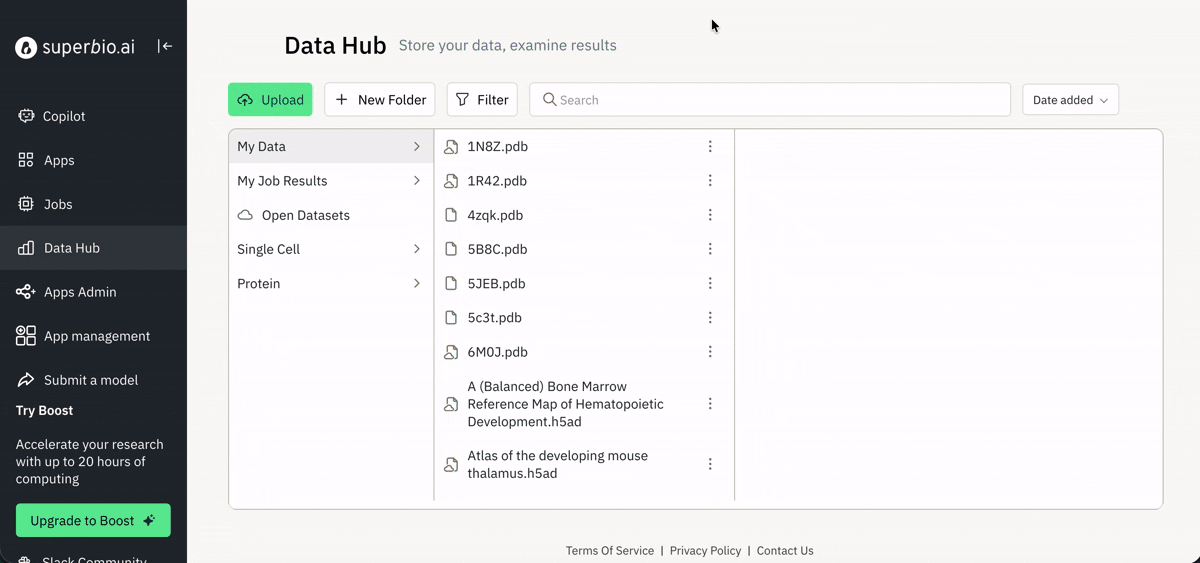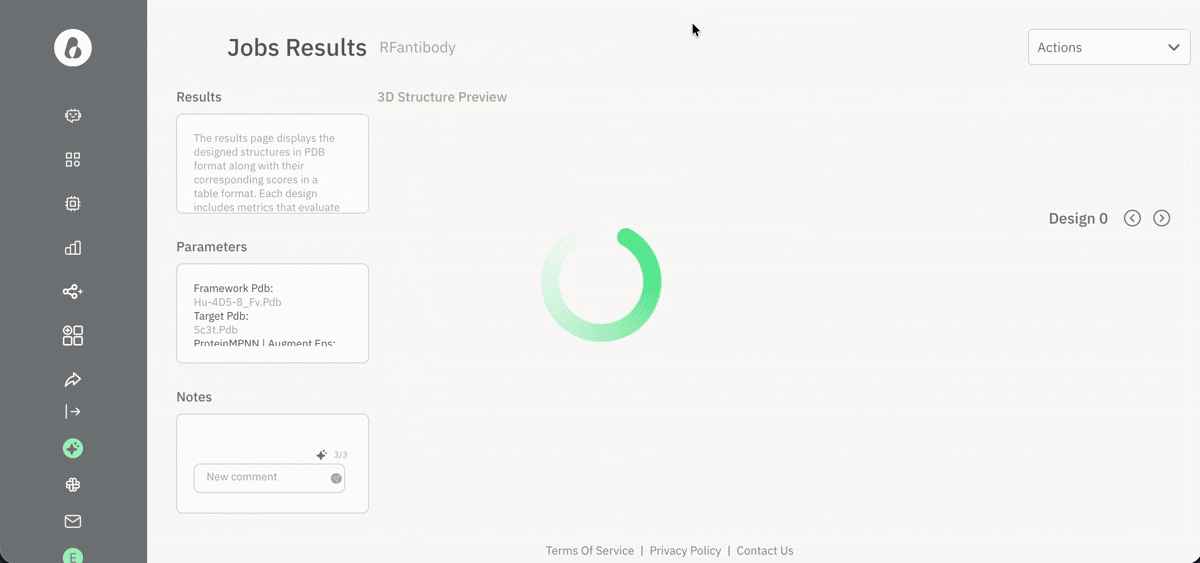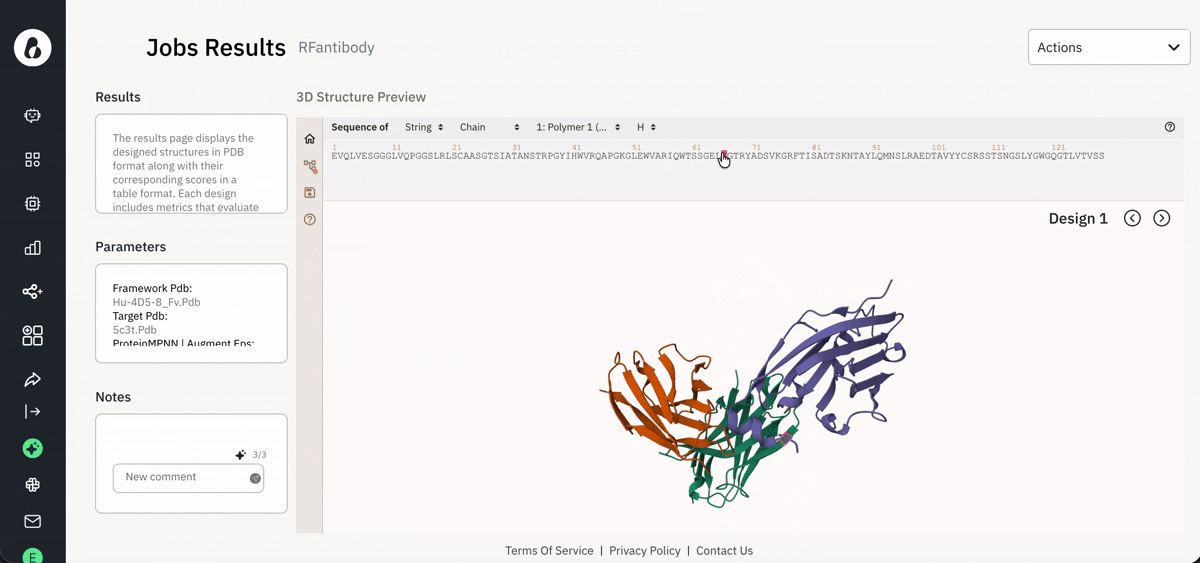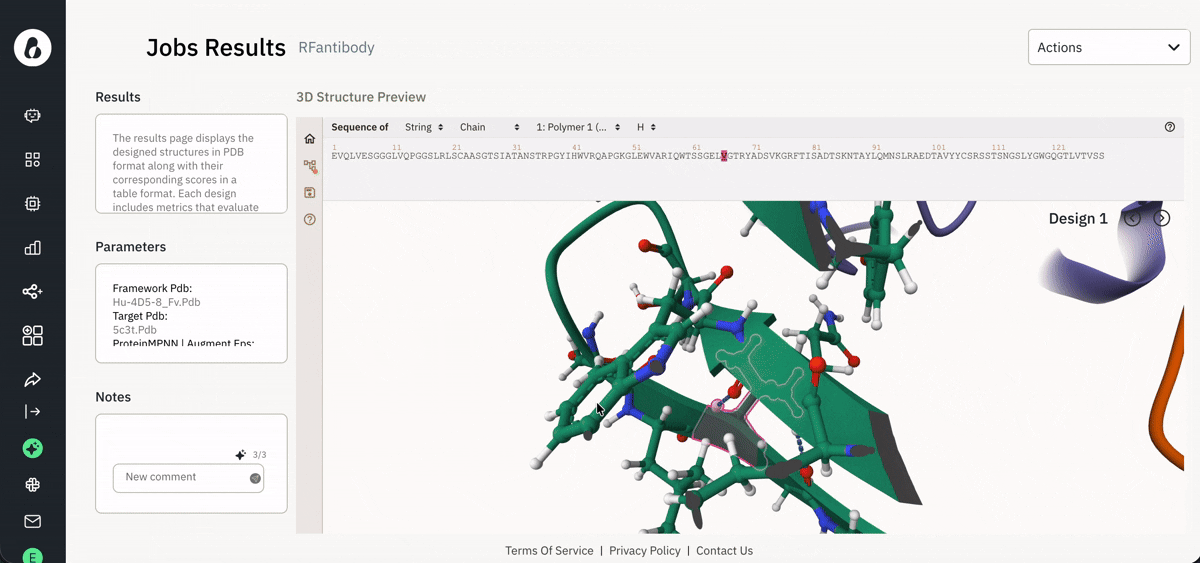
Exploring agent-generated data via the Jobs page.
Without further ado, try out Superbio’s Virtual Lab here: https://app.superbio.ai/
Have feedback on this feature, or want to share results? Just hit reply.
Until next time,
The Superbio Team 🩷